SEOUL, April 27 (AJP) - A research team led by Professor Sukjoon Yoon at Sookmyung Women's University in South Korea has created an ontology-based framework to quantitatively interpret gene-based biological data, the prominent university said Monday.
Biological data has historically been difficult for artificial intelligence to interpret because it lacks the clear contextual relationships found in standard text. To address this, the team used biological ontologies—systems that structure concepts for computer processing—to help AI analyze genetic information more effectively.
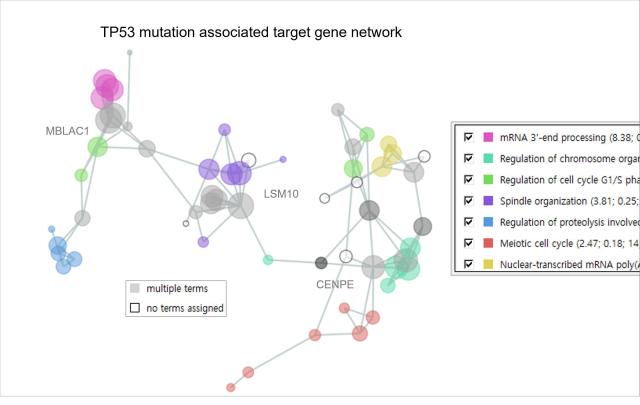
The researchers developed an algorithm called NetCrafter that measures how much functional or disease-related context-specific genes share. It converts these relationships into numerical networks, allowing for a more precise analysis of complex interactions within multi-omics data.
The team is currently working with CBiS Inc., a startup founded at the university, to build AI models that can automate the study of drug targets, biomarkers, and biological mechanisms. The NetCrafter technology is already available globally through the Q-omics platform, and the university is currently pursuing a patent for the system.
"This research will contribute to the realization of biological intelligence that understands data more deeply, moving beyond large language models," Professor Yoon said.
Professor Hohsuk Noh at Sookmyung Women's University's Department of Statistics and Eunah Chung, a researcher at the Research Institute of Women's Health, contributed to the study, alongside Dr. Jaemoon Shin from the Life Science Data Center in Japan. The findings appeared in the journal Briefings in Bioinformatics.
(Reference Information)
Journal/Source: Briefings in Bioinformatics (IF 7.7, JCR top 2.78 percent)
Title: NetCrafter: ontology-derived gene network modeling and functional interpretation
Link/DOI: https://doi.org/10.1093/bib/bbaf631.024
Copyright ⓒ Aju Press All rights reserved.